EEGLAB Day Slides
EEGLAB
- It offers a GUI to get you started
- Widely used in the EEG field
- It provides numerous techniques to apply to the data
BUT…
The simplicity hides the technicality and nuances of the analysis you apply to your data!
We hope that the first part of the day gave you useful knowledge to understand what you are doing while using EEGLAB GUI (or any other software)
Preprocessing (simplified)
- Load data
- Downsample
- Filter
- Change channel location
- Remove bad data
- Interpolate channels
- Rereference to average
- ICA
- Epoching
Load data
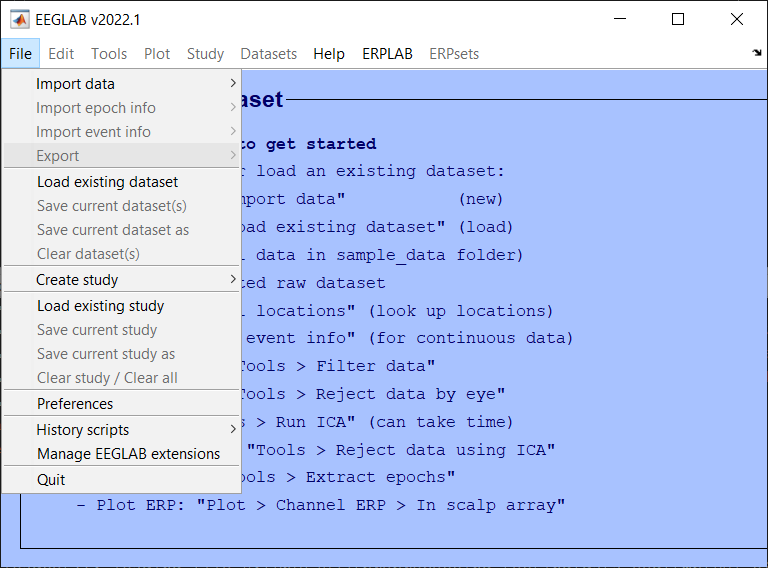
Load data

Downsample
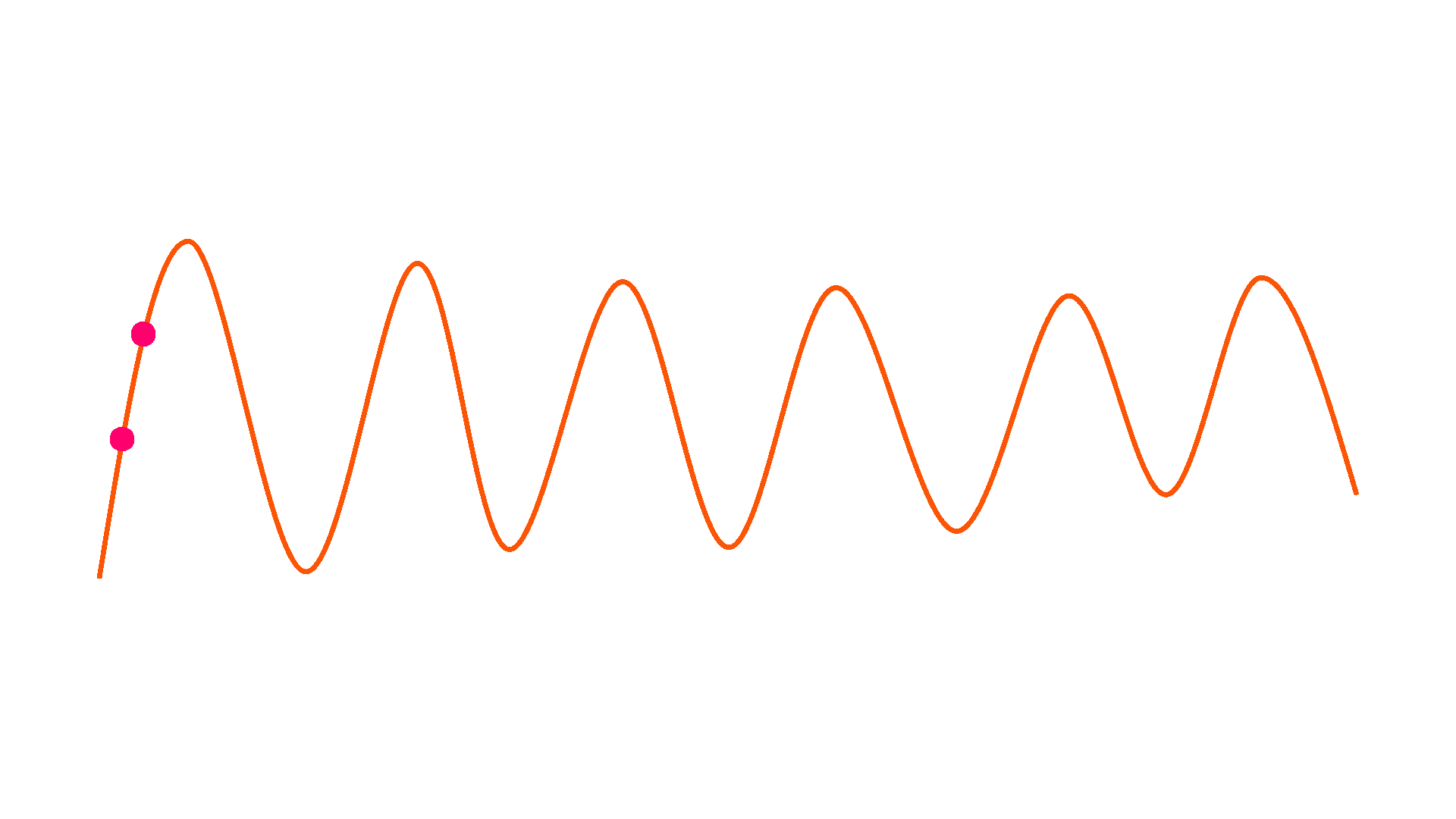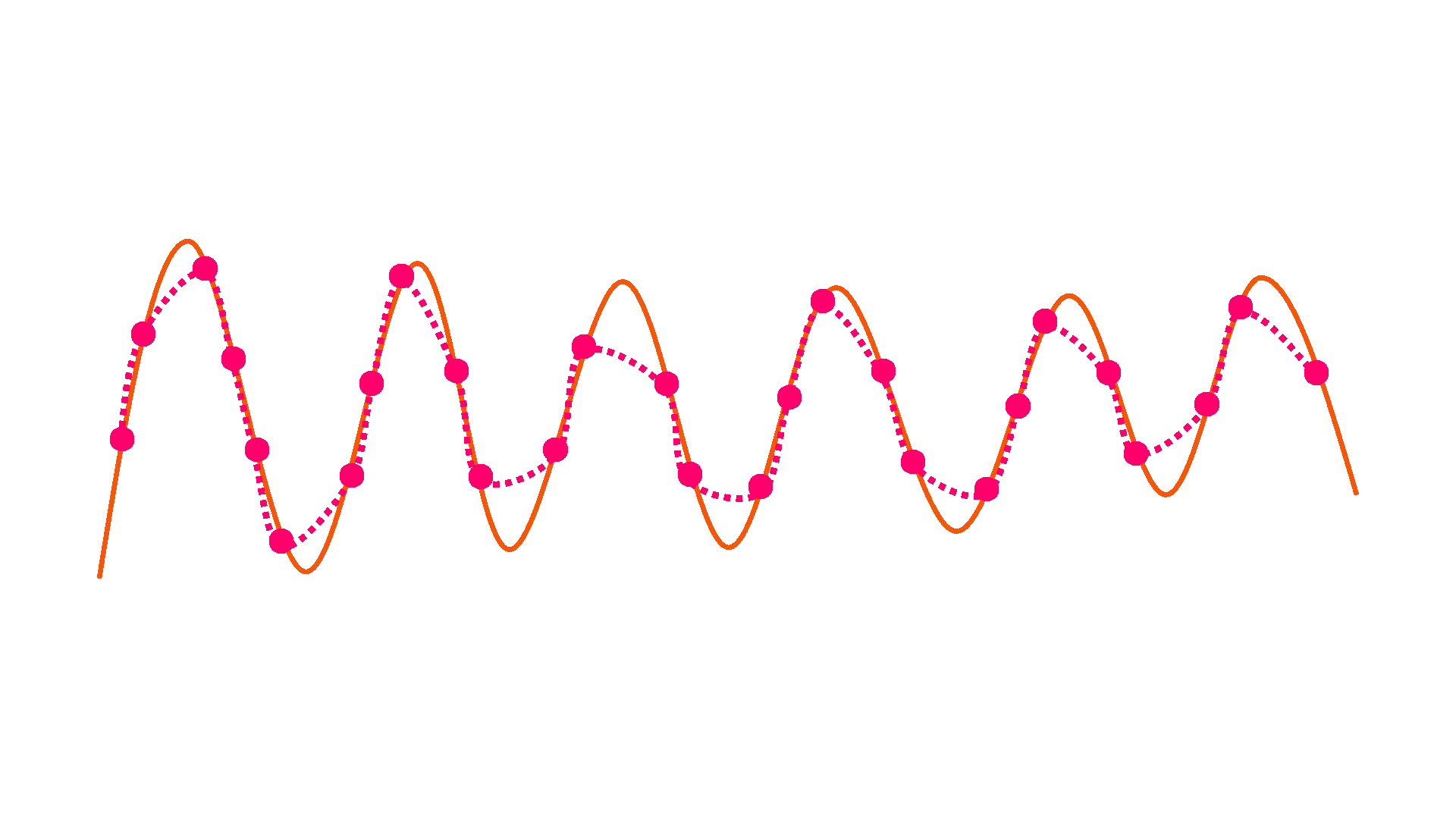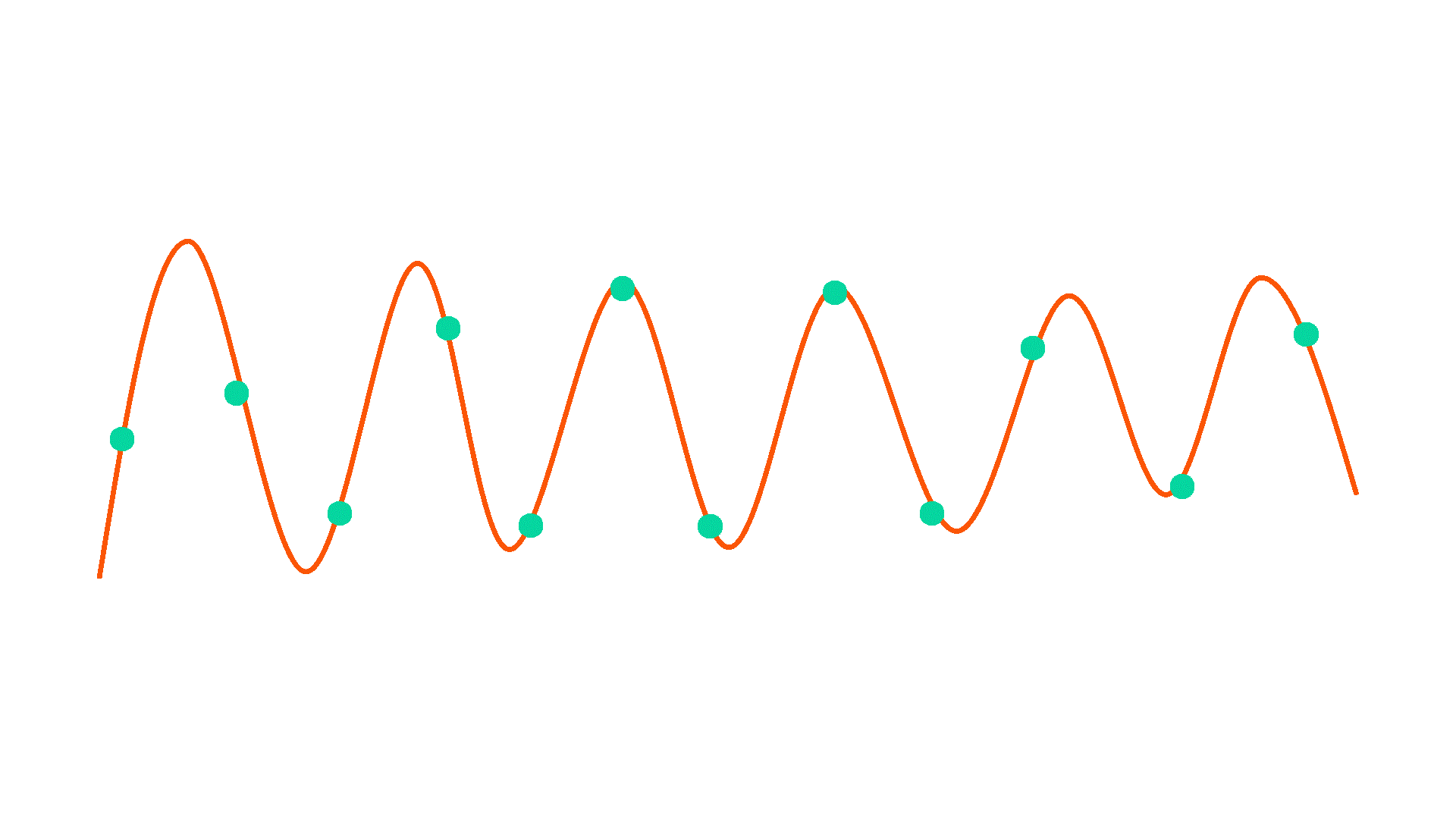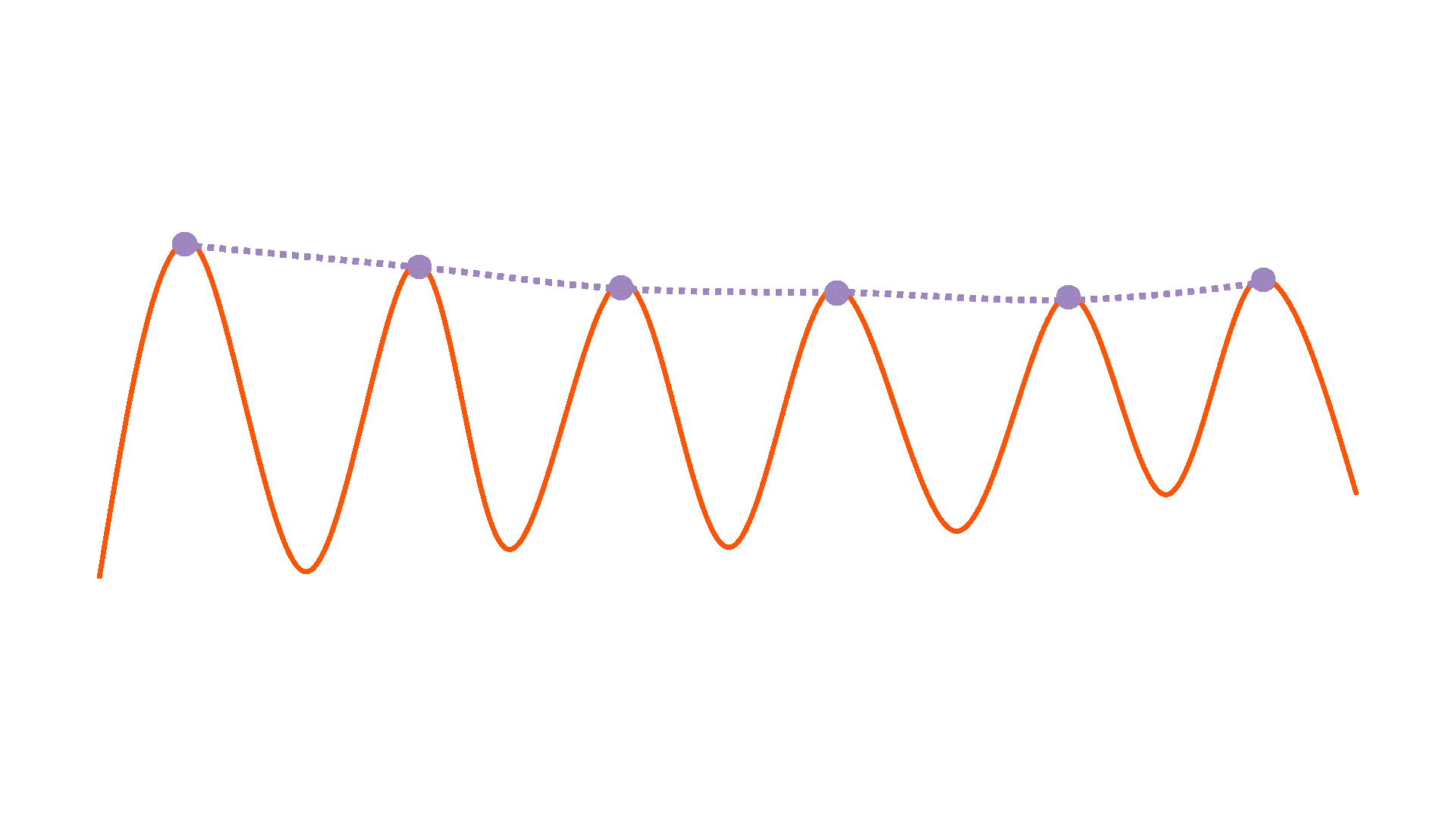
Downsample
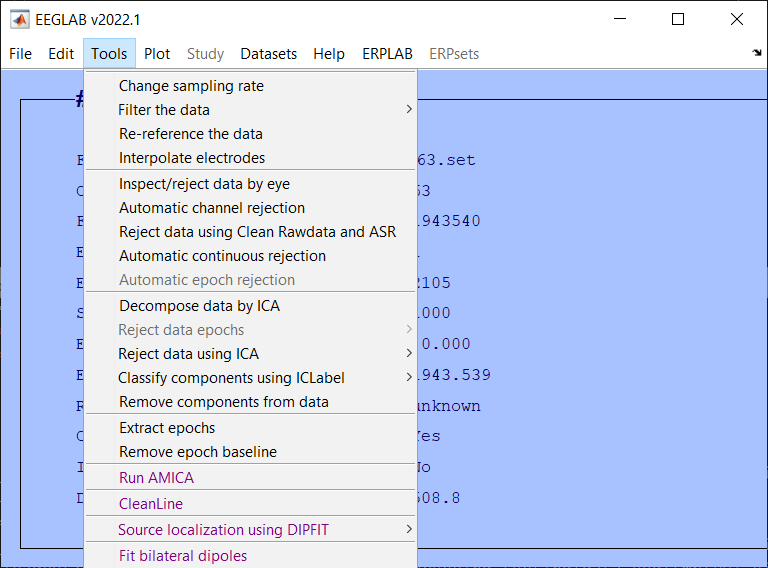
Downsample

Filter

Filter

Filter

Channel Location
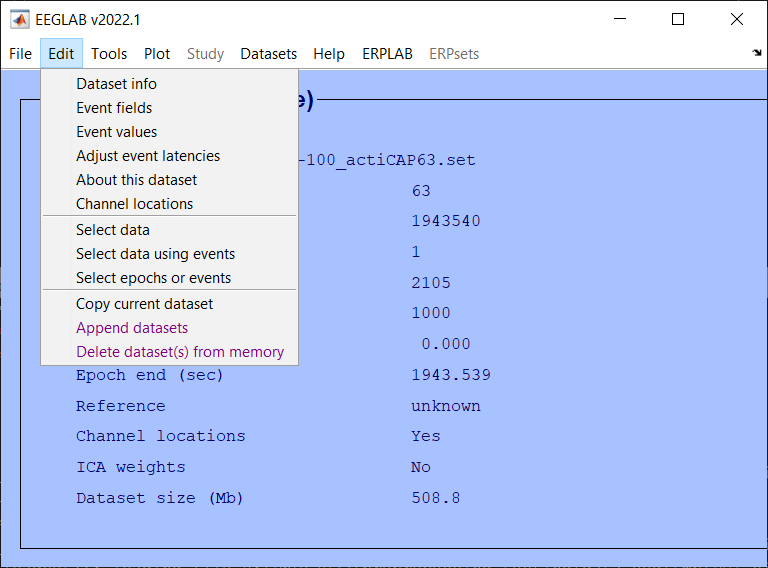
Channel Location

Remove bad channels

Remove bad channels

Remove bad channels

Interpolate removed channels

Interpolate removed channels

Interpolate removed channels

Rereference

ICA
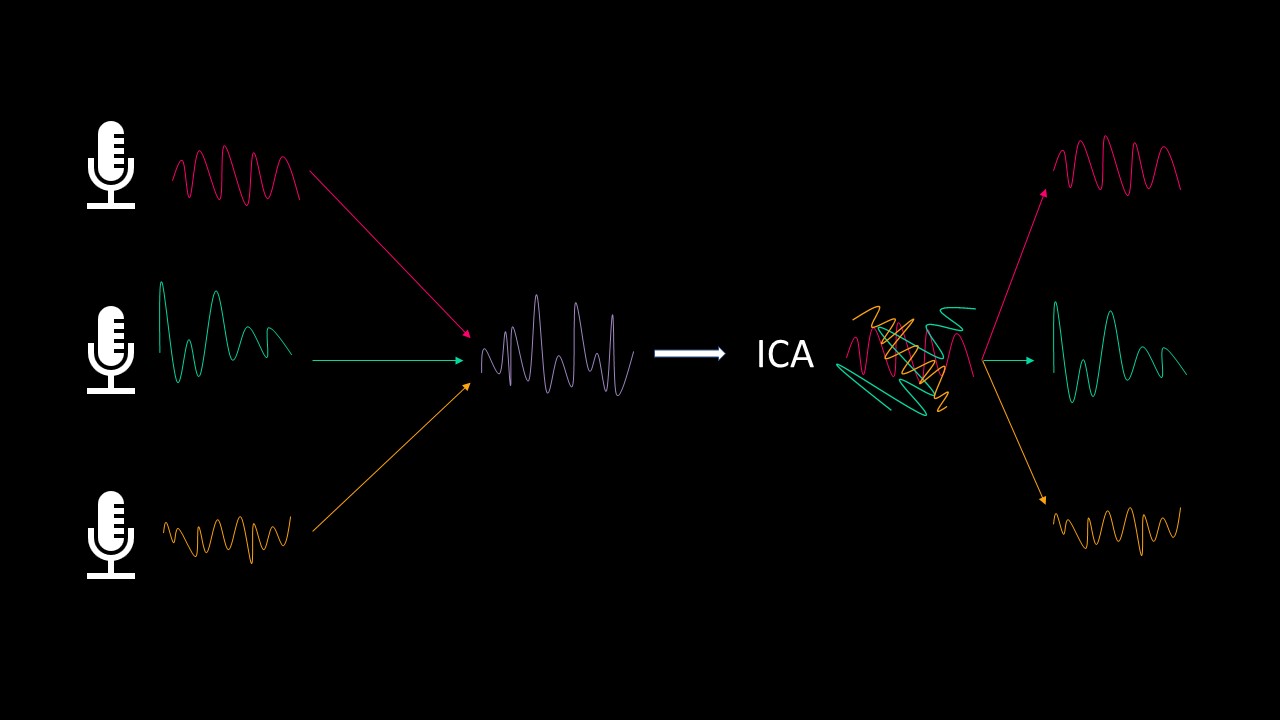
ICA
Before we run it
ICA
Got to lines 610-625 (at the time of writing)
function tmprank2 = getrank(tmpdata, pca_opt)
tmprank = rank(tmpdata);
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%Here: alternate computation of the rank by Sven Hoffman
%tmprank = rank(tmpdata(:,1:min(3000, size(tmpdata,2)))); old code
covarianceMatrix = cov(tmpdata', 1);
[~, D] = eig (covarianceMatrix);
rankTolerance = 1e-7;
tmprank2=sum (diag (D) > rankTolerance);
if tmprank ~= tmprank2
if nargin >= 2 && pca_opt ~= 0
fprintf('Warning: fixing rank computation inconsistency (%d vs %d) most likely because running under Linux 64-bit Matlab\n', tmprank, tmprank2);
end
tmprank2 = min(tmprank, tmprank2);
endICA
Got to lines 610-625 (at the time of writing)
function tmprank2 = getrank(tmpdata, pca_opt)
tmprank = rank(tmpdata);
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%Here: alternate computation of the rank by Sven Hoffman
%tmprank = rank(tmpdata(:,1:min(3000, size(tmpdata,2)))); old code
covarianceMatrix = cov(tmpdata', 1);
[~, D] = eig (covarianceMatrix);
rankTolerance = 1e-7;
tmprank2=sum (diag (D) > rankTolerance);
if tmprank ~= tmprank2
if nargin >= 2 && pca_opt ~= 0
fprintf('Warning: fixing rank computation inconsistency (%d vs %d) most likely because running under Linux 64-bit Matlab\n', tmprank, tmprank2);
end
tmprank2 = min(tmprank, tmprank2);
endICA
function tmprank2 = getrank(tmpdata, pca_opt)
tmprank = rank(tmpdata);
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%Here: alternate computation of the rank by Sven Hoffman
%tmprank = rank(tmpdata(:,1:min(3000, size(tmpdata,2)))); old code
covarianceMatrix = cov(tmpdata', 1);
[~, D] = eig (covarianceMatrix);
rankTolerance = 1e-7;
tmprank2=sum (diag (D) > rankTolerance);
if tmprank ~= tmprank2
if nargin >= 2 && pca_opt ~= 0
fprintf('Warning: fixing rank computation inconsistency (%d vs %d) most likely because running under Linux 64-bit Matlab\n', tmprank, tmprank2);
end
#tmprank2 = min(tmprank, tmprank2);
tmprank2 = max(tmprank, tmprank2);
endICA
- Please remember if you have applied this change
- Check that it’s doing what you expect
- Report that you have done this for replicability
ICA

ICA

ICA

ICA
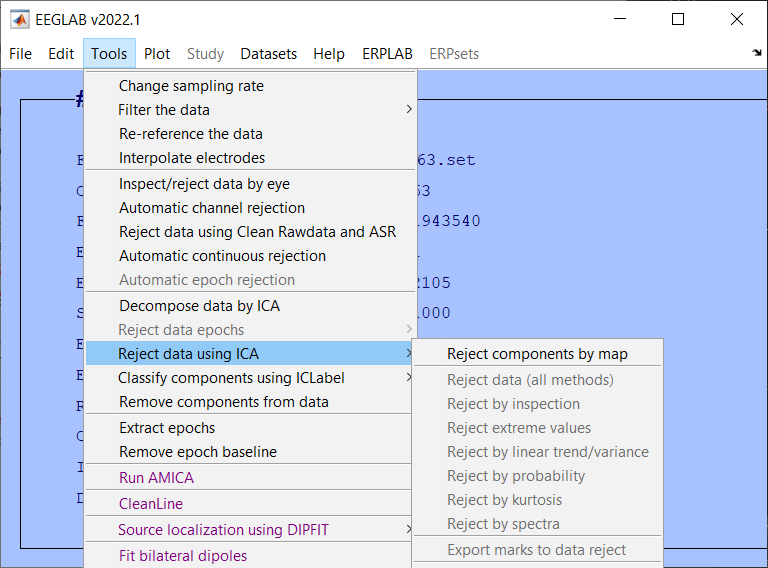
ICA

Epochs
